Software at CIS : BrainWorks: Tutorials: Segmentation
Tutorials | Modules | Change Log
- On the left column, click on the desired image so that it is highlighted in green. Select Image->Segment from the menu.
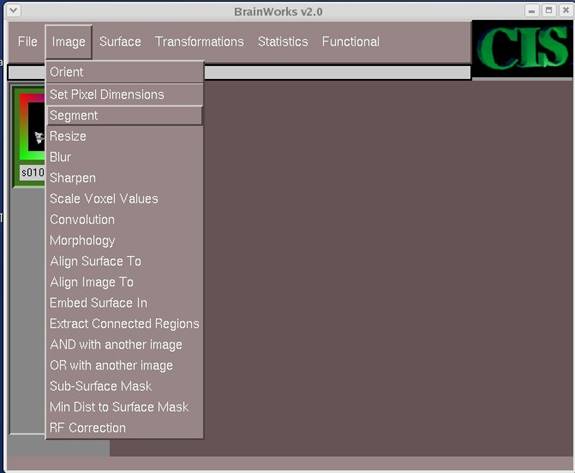
- Click OK on the pop-up screens except the last one. When asked to choose on fully automatic mode, click No.
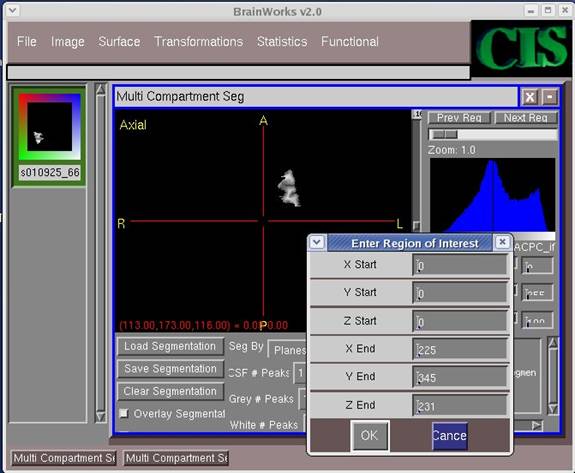
- Click on the "Seg By" field area and select "Cubes."
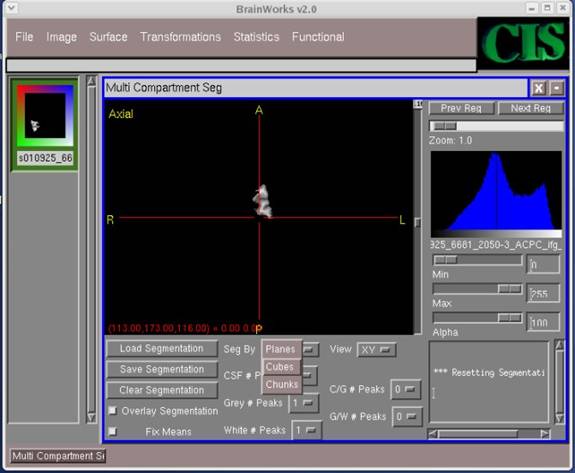
- Click OK on the pop-up screen to continue.
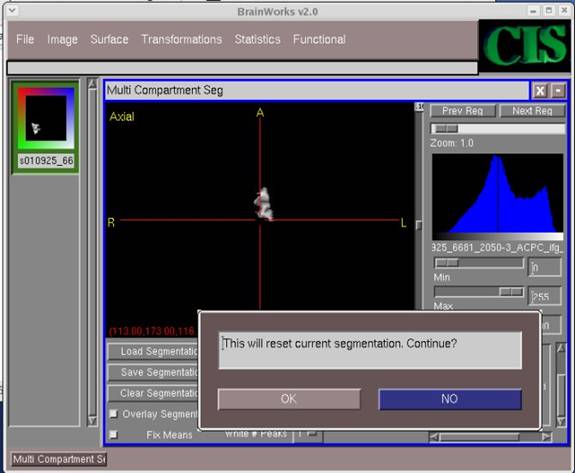
- Continue clicking OK on the following screens that pop-up (which define bounds for the image, etc.) until you reach the screen which asks the question, "Use Fully Automatic Mode?" For this question, select NO.
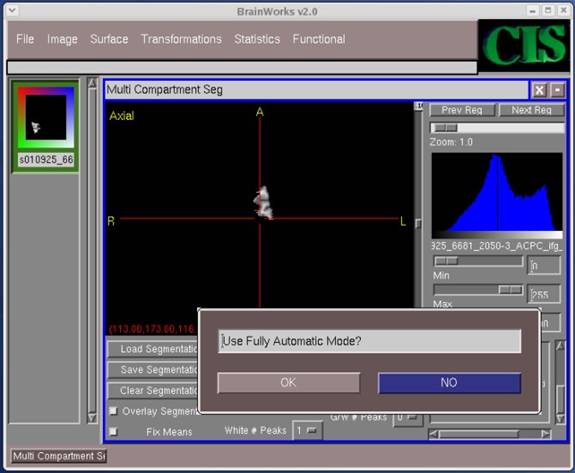
- For AKM or classical Bayesian, we want to segment our image into three distinct categories, namely gray matter (GM), white matter (WM), and CSF. For Neyman-Pearson recalibration, we add CSF/GM and GM/WM. We want to select the number of peaks for each of these categories, which could be 1, 2, etc. For example, the values for the indicated fields could be as follows:
AKM: CSF # Peaks -- 2 C/G # Peaks -- 0 Gray # Peaks -- 2 G/W # Peaks -- 0 White # Peaks -- 2 Bayesian: CSF # Peaks -- 1 C/G # Peaks -- 0 Gray # Peaks -- 1 G/W # Peaks -- 0 White # Peaks -- 1 Bayesian (Neyman-Pearson recalibration): CSF # Peaks -- 1 C/G # Peaks -- 1 Gray # Peaks -- 1 G/W # Peaks -- 1 White # Peaks -- 1
Select these values in the appropriate fields.
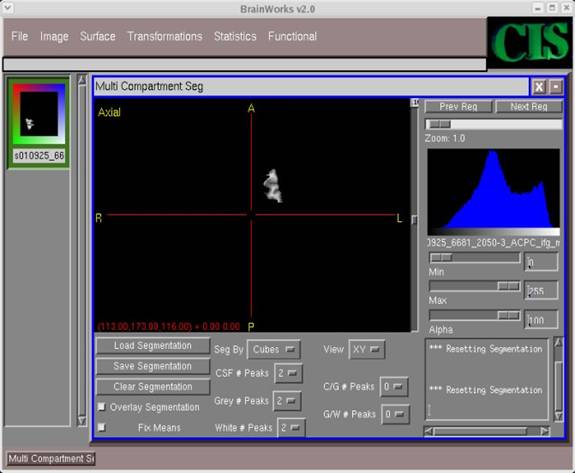
- In the window with the image, click around with the cursor so that the image is centered. Then use the horizontal Zoom bar in the top-right corner to zoom in and out and obtain the desired image display.
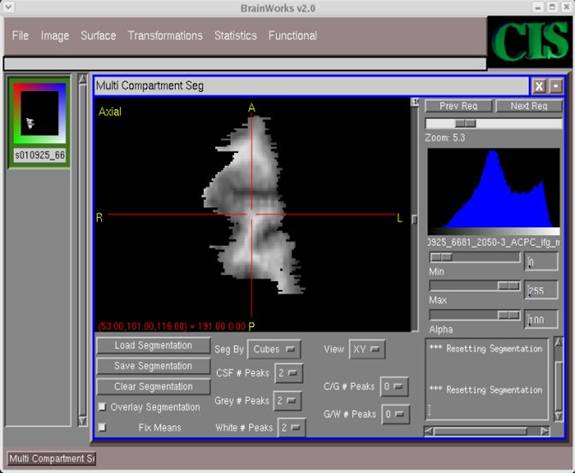
- To execute the segmentation, locate the histogram window on the right. Since we have selected 6 total peaks in this example, we will need to click 6 times in this window. Each click should correspond to a rough estimate for where the peak value(s) occurs in each category (CSF, GM, WM). For each click, a vertical green line will appear.
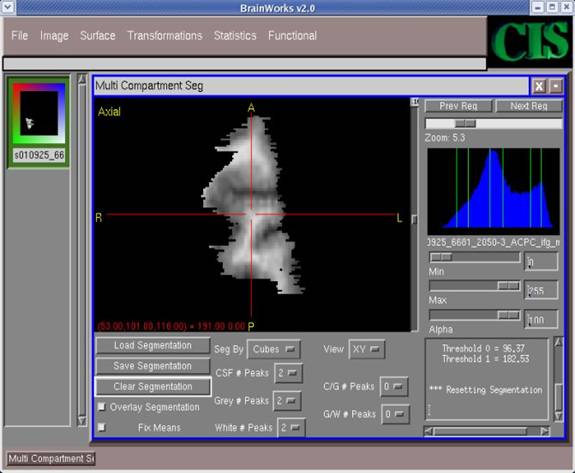
- After a short delay, the segmented image will appear in the main window. In addition, the histogram will have Gaussian curves which estimate the histogram. In the bottom right corner, threshold frequencies are calculated to establish the borderline frequencies between the different categories. The image is segmented such that
CSF < Threshold 0 < GM < Threshold 1 < WM.
CSF will appear red (with value 25 by default), GM blue (with value 75 by default), and WM white (with value 125 by default). Partial volumes CSF/GM and GM/WM have default values of 50 and 100 respectively.
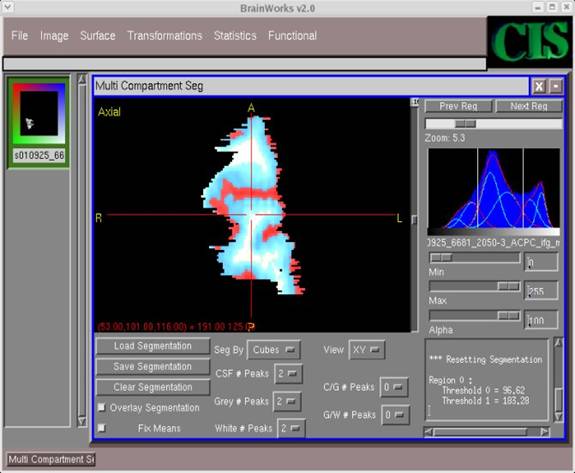
- Ideally, when clicking in the histogram window to estimate peaks (as in step 15), the same threshold frequencies should be produced regardless of where you click. However, this is hardly ever the case. Therefore, you should repeat step 15 several times by clicking in different areas of the histogram and monitor how the threshold frequencies change. Here is an example of a segmentation where the CSF (in red) is clearly overestimated.
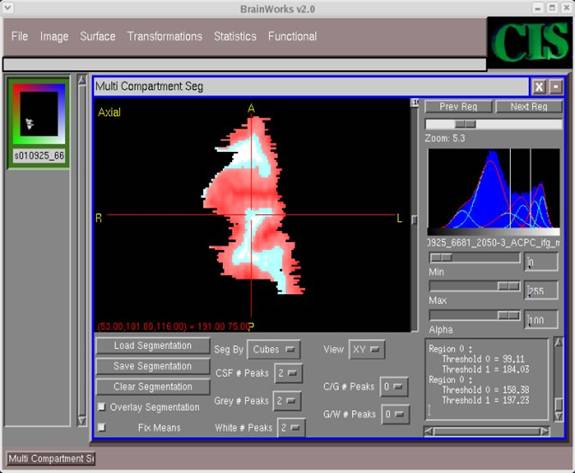
- As stated in Step 6, we can specify any number of peaks for each category for the segmentation. In this example, we have selected the following
CSF # Peaks -- 2 C/G # Peaks -- 0 Gray # Peaks -- 1 G/W # Peaks -- 0 White # Peaks -- 1
Clearly, this combination of peak values does not produce a desirable segmentation. To find the best segmentation, it is just a matter of experimenting with different combinations of peaks, and seeing how the threshold frequencies vary for each combination.
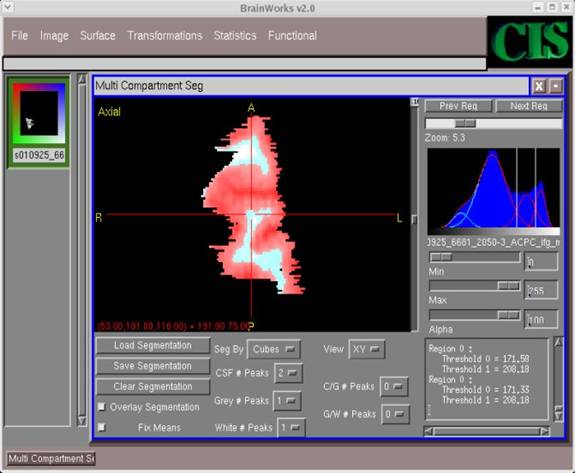
- After several different segmentation attempts, you should be able to decide on one which bests segments the given image. (Use the darker and lighter areas in the image as a guide, e.g. darker areas are closer to CSF, medium areas GM, and lighter areas WM.) After obtaining the best segmentation, click Save Segmentation.
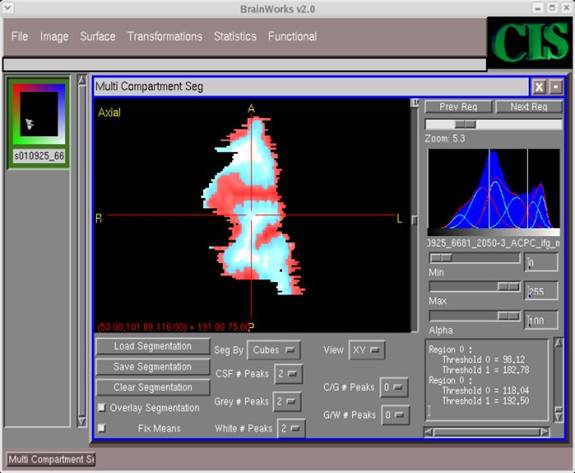
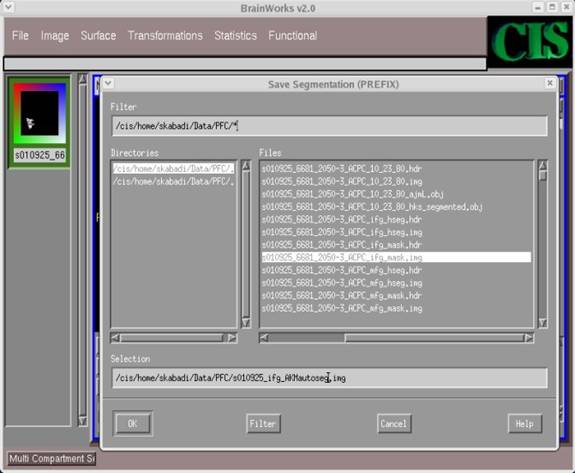
- Once again, navigate through the directories to the appropriate one, and type in an appropriate file name for the auto-segmentation file in the Selection field (along with extension .img). Then click OK.
Last Modified: Monday, 25th April, 2011 @ 11:19am


